Enhancing Early Detection of Esophageal Squamous Cell Carcinoma through Shallow Whole-Genome Sequencing and Non-Endoscopic Sponge Sampling
Authors
Aisha Yusuf1, Maryla H. Turkot2, Shalini Malhotra3, Maria O’Donovan3, Massimiliano di Pietro1, Nastazja D. Pilonis2, Karol Nowicki-Osuch4, Andrzej Mroz2,5, Malgorzata Lenarcik2,5, Michal Mikula6, Monika Wojtowicz-Popiel2,7, Massimiliano di Pietro1, Jaroslaw Regula2,7, Michal F. Kaminski2,7, Rebecca C. Fitzgerald*1, Wladyslaw Januszewicz*2,7
* joint senior co-authors
Affiliations
- Early Cancer Institute, University of Cambridge, United Kingdom
- Department of Gastroenterology, Hepatology and Clinical Oncology, Centre of Postgraduate Medical Education, Warsaw, Poland
- Department of Histopathology, Cambridge University Hospitals NHS Foundation Trust, Cambridge, UK
- Herbert and Florence Irving Institute for Cancer Dynamics, Columbia University, New York, USA
- Department of Pathomorphology, Centre of Postgraduate Medical Education, Warsaw, Poland
- Department of Genetics, Maria Sklodowska-Curie National Research Institute of Oncology, Warsaw, Poland
- Department of Gastroenterological Oncology, Maria Sklodowska-Curie National Research Institute of Oncology, Warsaw, Poland
Introduction and Aims
Oesophageal squamous cell carcinoma (OSCC) accounts for nearly half a million global annual cases of oesophageal cancer, often developing through detectable copy number abnormalities (CNAs). However, the expense and invasiveness of traditional endoscopy limit its use in monitoring at-risk groups. This study tested shallow whole-genome sequencing (sWGS) using a non-endoscopic sponge device, Cytosponge, to detect CNAs for the early detection of OSCC. We aimed to develop and validate a model for early detection of oesophageal dysplasia and neoplasia, improving prognosis through timely intervention.
Methods
We collected oesophageal specimens from 178 individuals using the Cytosponge device: 24 with OSCC, 61 high-risk, and 93 healthy controls with dyspeptic symptoms. The samples underwent 1.5-2x sWGS, H&E staining, and p53 immunohistochemistry and were analysed by GI pathologists blinded to the reference endoscopic diagnoses. Penalised regression identified significant CNAs linked to early neoplastic changes. We refined a logistic model for predicting these changes in Cytosponge-derived specimens and validated it against sWGS from microdissected endoscopic submucosal dissected (ESD) samples.
Results
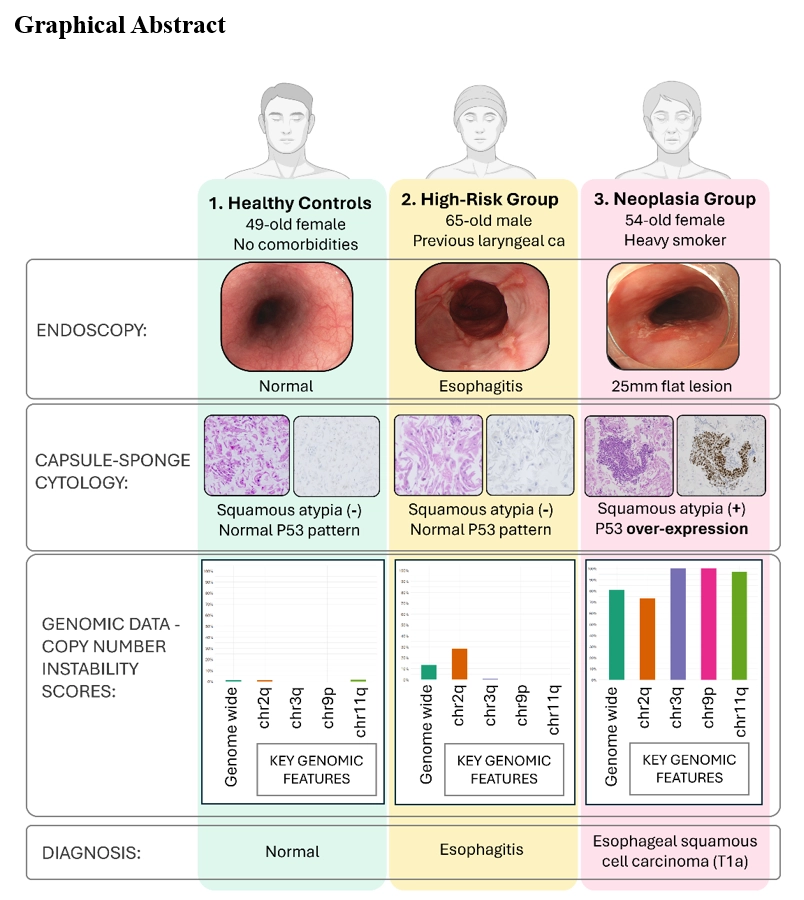
We identified significant genomic markers for oesophageal squamous dysplasia and early neoplasia, including CNAs on chr2q, chr3q, chr9p, and chr11q, and the Genome-Wide Instability Score (GWIS). Our logistic regression model, incorporating these biomarkers, achieved an AUROC of 0.947, with a sensitivity and specificity of 89% and 88.2%, respectively, surpassing a model based solely on pathological data (AUROC 0.880).
Combining genomic and pathological data further improved the AUROC to 0.969, with the sensitivity and specificity reaching 95.9% and 90.5%, respectively. Further validation of the genomic model against microdissected ESD samples yielded an AUROC of 0.879, improving to 0.905 after excluding low-grade dysplasia (LGD), highlighting diagnostic variability in LGD classification.
Conclusion
The integration of sWGS with non-endoscopic sponge sampling has the potential for early OSCC detection, especially in high-risk populations where traditional endoscopy is impractical.