Plasma Metabolomics and Machine Learning Identify Biomarkers of Gastric Cancer
Authors
Juan Zhu1#, Yida Huang2,3#, Bin Liu1,4#, Xue Li1, Li Yuan5, Le Wang1, Kun Qian2,3, Yingying Mao4, Lingbin Du1,6*, Xiangdong Cheng5,7*, Hongbing Shen8*
Affiliations
- Department of Cancer Prevention, Zhejiang Cancer Hospital, Hangzhou Institute of Medicine (HIM), Chinese Academy of Sciences, Hangzhou, China
- State Key Laboratory of Systems Medicine for Cancer, School of Biomedical Engineering, Institute of Medical Robotics and Med-X Research Institute, Shanghai Jiao Tong University, Shanghai 200030, China
- Division of Cardiology, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200127, China
- Department of Epidemiology, School of Public Health, Zhejiang Chinese Medical University, Hangzhou 310053, China
- Department of Gastric Surgery, Zhejiang Cancer Hospital, Hangzhou 310022, China
- School of Public Health and Management, Wenzhou Medical University, Wenzhou, Zhejiang, 325035, China.
- Key Laboratory of Prevention, Diagnosis and Therapy of Upper Gastrointestinal Cancer of Zhejiang Province, Hangzhou, China
- Department of Epidemiology, Center for Global Health, School of Public Health, Nanjing Medical University, Nanjing 211166, China
Introduction
Current detection methods for Gastric cancer(GC) mainly rely on gastroscopy, limited by a relatively low uptake. Metabolic biomarkers are expected to decode the phenotype of GC and offer promise for GC diagnosis. This study aims to identify metabolic markers and develop a diagnostic model for GC using a multi-step strategy.
Methods
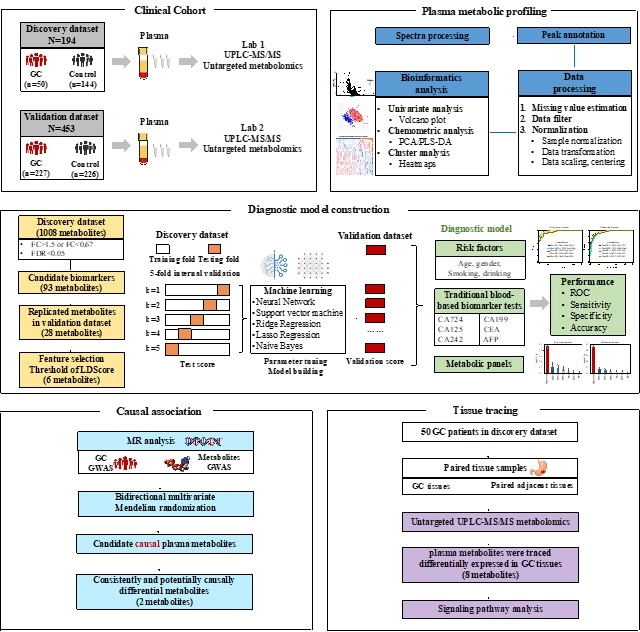
We conducted a two-phase case-control study involving 647 participants, comprising 277 GC cases and 370 non-GC individuals. Through UPLC-MS platform, candidate differential metabolites were initially identified using fold change and false discovery rate criteria, followed by optimization of LRScore-based selection based on AUC calculations with varying thresholds of LRScore. Diagnostic modes were developed in the discovery and verification phases using machine learning algorithms, including neural network, support vector machine, ridge regression, lasso regression and Naive Bayes. Bidirectional two-sample Mendelian randomization(MR) analysis examined the causal effect of metabolic biomarkers on GC. Additionally, tumour specificity of plasma markers was then confirmed by comparison with tumor-adjacent non-malignant paired tissue.
Results
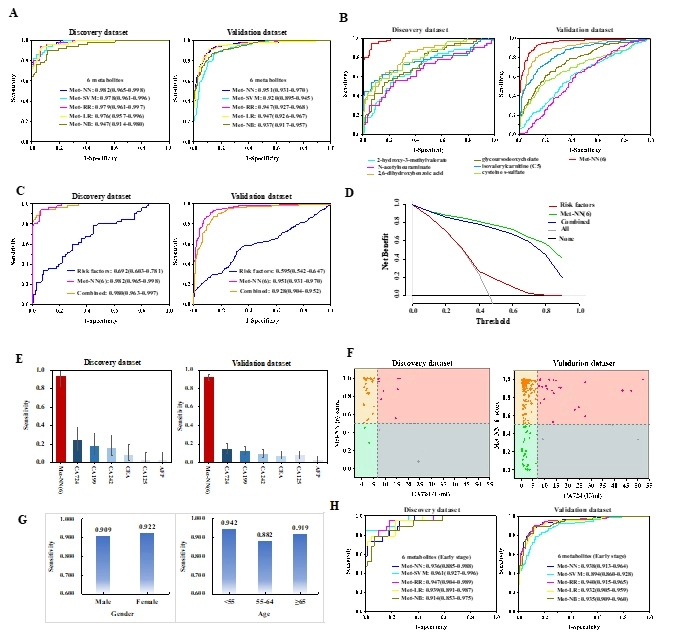
Twenty-eight replicated plasma metabolites were identified in the discovery and validation dataset. Of these, six metabolic features were selected to construct a metabolic panel, which exhibited excellent diagnostic performance, achieving AUCs ranging from 0.947-0.982 in the discovery dataset and 0.920-0.951 in the independent external verification dataset through machine learning modes. The diagnostic sensitivity of the biomarker panel(0.900-0.940) significantly outperformed traditional clinical protein biomarker tests(0.020-0.240). Moreover, the panel exhibited desirable performance in early GC diagnosis, recording AUCs of 0.914 to 0.961 in the discovery set and 0.894 to 0.940 in the validation set. Eight metabolites were traced differentially expressed in GC and paired adjacent tissues, and two causal plasma metabolites were identified in MR analysis (2-hydroxy-3-methylvalerate, isovalerylcarnitine (C5)).
Conclusion
This study identifies promising metabolic biomarkers for GC diagnosis and develops a validated diagnostic model, providing insights into early detection of GC.